Characterizing Cellular Fatty Acid Methyl Ester (FAME) Profiles to Identify Bacteria Using Gas Chromatography
Dr. Radomír Čabala is the Head of the Toxicology Department at the General University Hospital in Prague, Czech Republic. His current research interests include the characterization of solute-solvent interactions by means of inverse GC; development, optimization, and testing of microextraction techniques; and comprehensive GC.
Practical, accurate methods for bacterial identification are of great interest in fields ranging from clinical diagnostics to food safety. Traditional methods, based on staining or growth, often are not specific enough to identify closely related species, which has led to the development of molecular methods based on DNA sequence or protein expression. Today, ELISA and other immunochemical techniques are commonly used, but gas chromatography can also be an effective approach. When paired with highly inert, low-bleed columns of appropriate selectivity, GC is one of the most efficient analytical separation methods available. The fatty acid methyl ester (FAME) analyses shown here illustrate the potential of GC-TOFMS for distinguishing clinically relevant species and identifying low-level compounds in these complex biological samples.
In the first example, the fatty acid composition of cell membranes from two Gram-negative bacteria, Bordetella bronchiseptica and Bordetella pertussis, were compared. B. bronchiseptica rarely infects humans, but it can cause infectious bronchitis and is resistant to macrolides and cephalosporins. B. pertussis is the dangerous agent of pertussis (whooping cough) in humans and produces several toxins. These species have a very close evolutionary relationship, which can be seen in their genetic similarity and cellular fatty acid profiles. To determine if these species could be distinguished based on FAME analysis, samples were first prepared from bacterial cell wall lysate by transesterification with sodium methanolate. Samples were then analyzed on a DANI Master GC-TOFMS equipped with an Rxi-17Sil MS column. This column was chosen based on its selectivity for the cellular fatty acids. Even without optimizing the method for separation efficiency, good separation of the saturated acid methyl esters and their respective unsaturated counterparts was achieved (Figure 1). This separation also shows that B. bronchiseptica produces hexadecanoic and hexadecenoic acids at a different ratio than B. pertussis. In addition, 2-hexyl-cyclopropane-octanoic acid is present in the B. bronchiseptica sample, but not in the B. pertussis sample. Cell membrane lysates of B. bronchiseptica were also cultured at 24 °C and 37 °C to investigate the effect of temperature. It is commonly observed that at lower temperatures microorganisms increase the fraction of unsaturated acids in the cell membrane in order to maintain membrane fluidity. This effect is evident for the pairs of hexadecanoic/hexadecenoic and octadecanoic/octadecenoic acids (Figure 2).
Figure 1: Differences in FAME profiles in cell membrane lysates can be useful for bacterial identification.
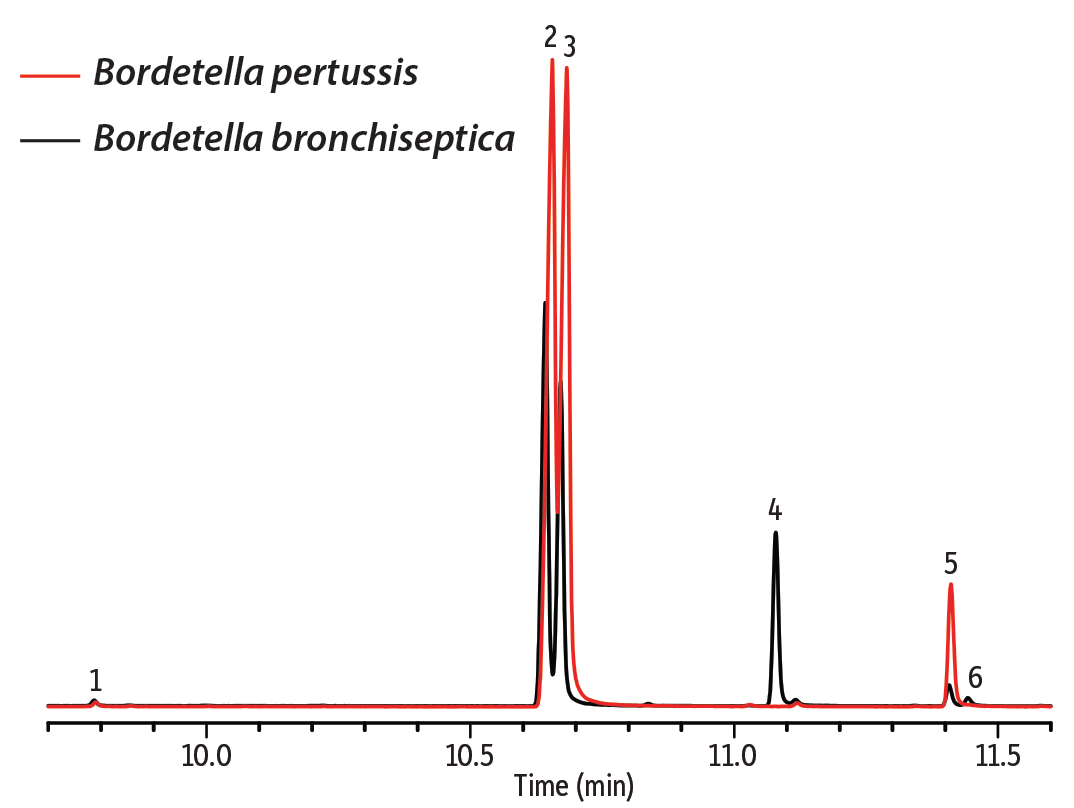
| Peaks | |
|---|---|
| 1. | Tetradecanoic acid methyl ester |
| 2. | Hexadecanoic acid methyl ester |
| 3. | Hexadecenoic acid methyl ester |
| 4. | 2-Hexyl-cyclopropane-octanoic acid methyl ester |
| 5. | Octadecanoic acid methyl ester |
| 6. | 9-Octadecenoic acid methyl ester |
| Column | Rxi-17Sil MS, 20 m, 0.15 mm ID, 0.15 µm (cat.# 43821) |
|---|---|
| Standard/Sample | FAMEs in bacterial cell membrane lysates |
| Injection | |
| Inj. Vol.: | 1 µL split (split ratio 50:1) |
| Inj. Temp.: | 250 °C |
| Oven | |
| Oven Temp.: | 70 °C (hold 2 min) to 250 °C at 25 °C/min (hold 10 min) |
| Carrier Gas | He, constant linear velocity |
| Linear Velocity: | 50 cm/sec |
| Detector | DANI Master GC-TOFMS @ 25 Hz |
|---|---|
| Acknowledgement | Data provided by Dr. Radomír Čabala, Head of the Toxicology Department at the General University Hospital in Prague, Czech Republic |
Figure 2: As expected, the ratio of unsaturated to saturated fatty acids in the cell membrane changes with temperature.
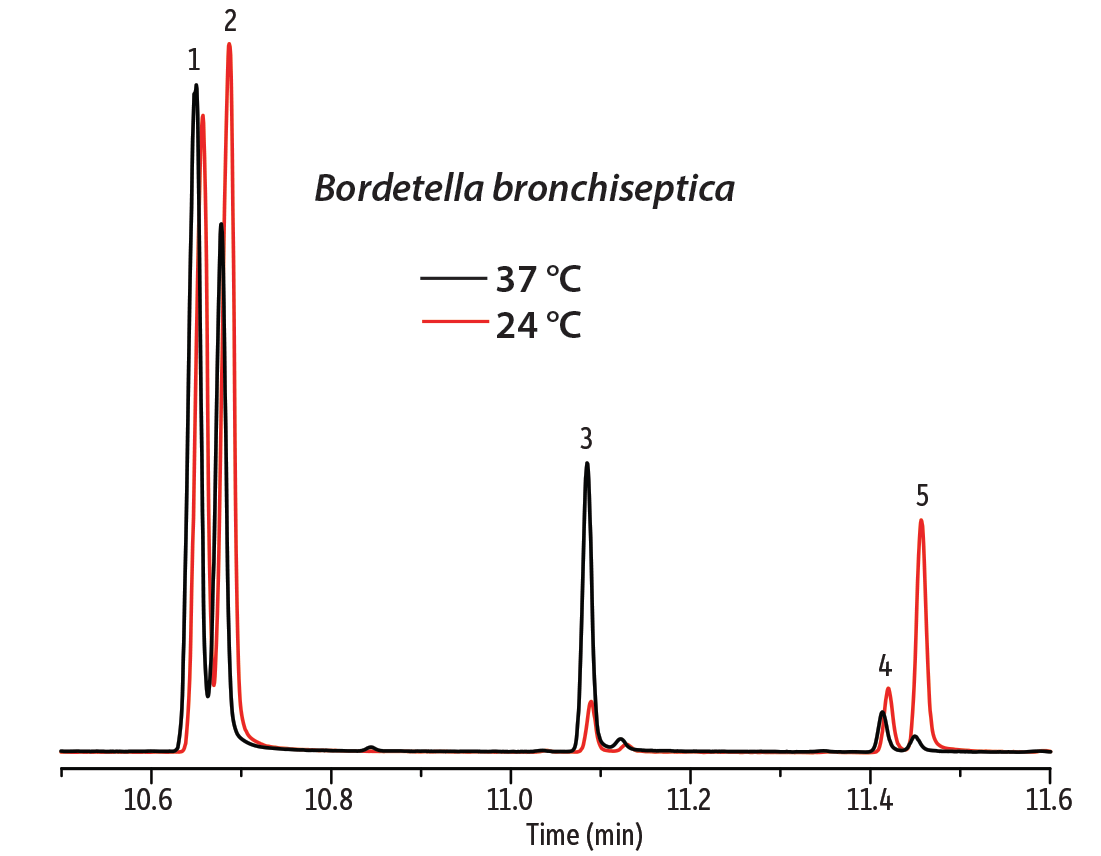
| Peaks | |
|---|---|
| 1. | Hexadecanoic acid methyl ester |
| 2. | Hexadecenoic acid methyl ester |
| 3. | 2-Hexyl-cyclopropane-octanoic acid methyl ester |
| 4. | Octadecanoic acid methyl ester |
| 5. | 9-Octadecenoic acid methyl ester |
See Figure 1 for instrument conditions.
In the second example, we investigated whether octadecanoic acid could be determined in phosphatidylcholin (lecithin) isolated from the cell membrane of the Gram-positive bacterium Bacillus subtilis. This bacterium produces a cyclopeptide surfactin that is one of the most potent natural antibiotics known. The problem arising here is the severe coelution of low levels of octadecanoic acid with the excess of unsaturated octadecenoic acids (Figure 3). Data were collected at 25 Hz, which was determined to be the optimum frequency, but octadecanoic acid was not chromatographically resolved. The signal processing feature of the TOF detector and the deconvolution option of the DANI Master Lab software allowed successful separation of the selected m/z signals and reliable compound identification of octadecanoic acid (match factor 835) based on the NIST 08 spectral database (Figure 4).
The examples presented here illustrate the potential utility of GC-TOFMS and an Rxi-17Sil MS column for bacterial identification through cellular fatty acid analysis. The efficiency and selectivity of the column and power of the instrument allow FAME profiles to be compared and a low-level coeluting compound (octadecanoic acid) to be distinguished.
Figure 3: Octadecanoic acid coelutes with the unsaturated octadecenoic acids and cannot be distinguished.
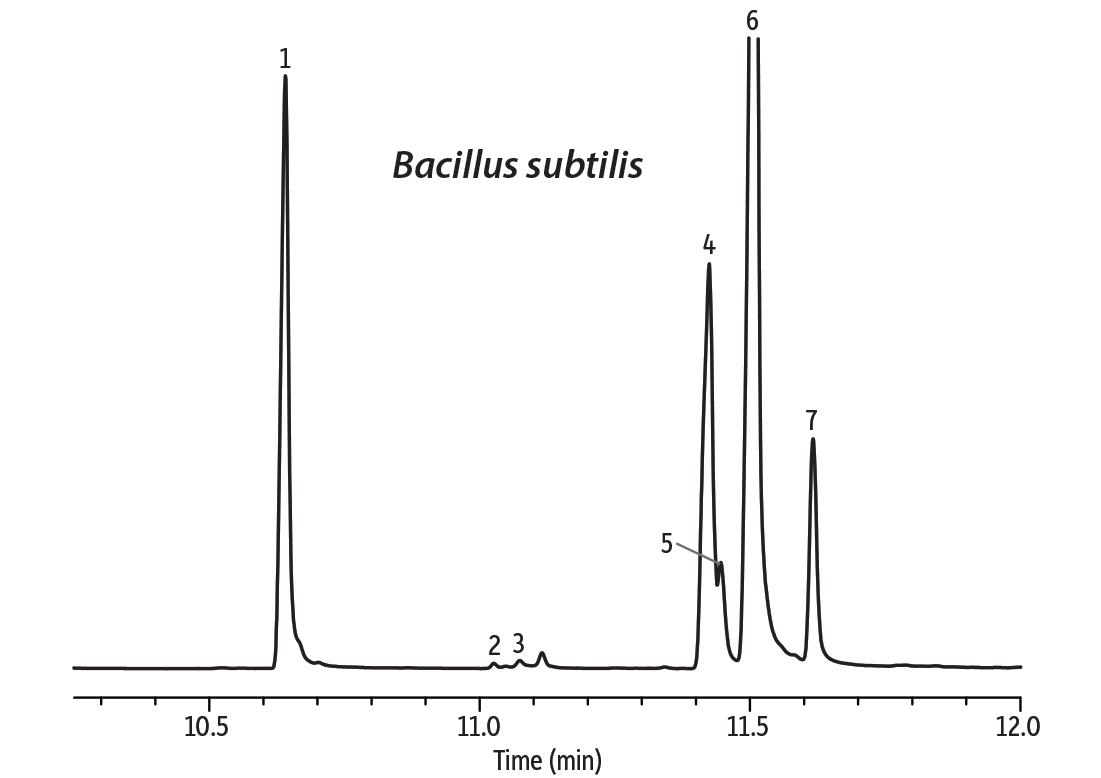
| Peaks | |
|---|---|
| 1. | Tetradecanoic acid methyl ester |
| 2. | 15-Methyl-hexadecanoic acid methyl ester |
| 3. | 2-Hexyl-cyclopropane-octanoic acid methyl ester |
| 4. | 9-Octadecenoic acid methyl ester |
| 5. | 11-Octadecenoic acid methyl ester |
| 6. | 9,12-Octadecadienoic acid methyl ester |
| 7. | 9,12,15-Octatrienoic acid methyl ester |
| Column | Rxi-17Sil MS, 20 m, 0.15 mm ID, 0.15 µm (cat.# 43821) |
|---|---|
| Standard/Sample | FAMEs in bacterial cell wall lecithin |
| Injection | |
| Inj. Vol.: | 1 µL split (split ratio 100:1) |
| Inj. Temp.: | 250 °C |
| Oven | |
| Oven Temp.: | 70 °C (hold 4 min) to 250 °C at 25 °C/min (hold 10 min) |
| Carrier Gas | He, constant linear velocity |
| Linear Velocity: | 50 cm/sec |
| Detector | DANI Master GC-TOFMS @ 25 Hz |
|---|---|
| Acknowledgement | Data provided by Dr. Radomír Čabala, Head of the Toxicology Department at the General University Hospital in Prague, Czech Republic |
Figure 4: TOFMS allows octadecanoic acid to be positively identified (match factor 835) even in the presence of coeluting peaks.
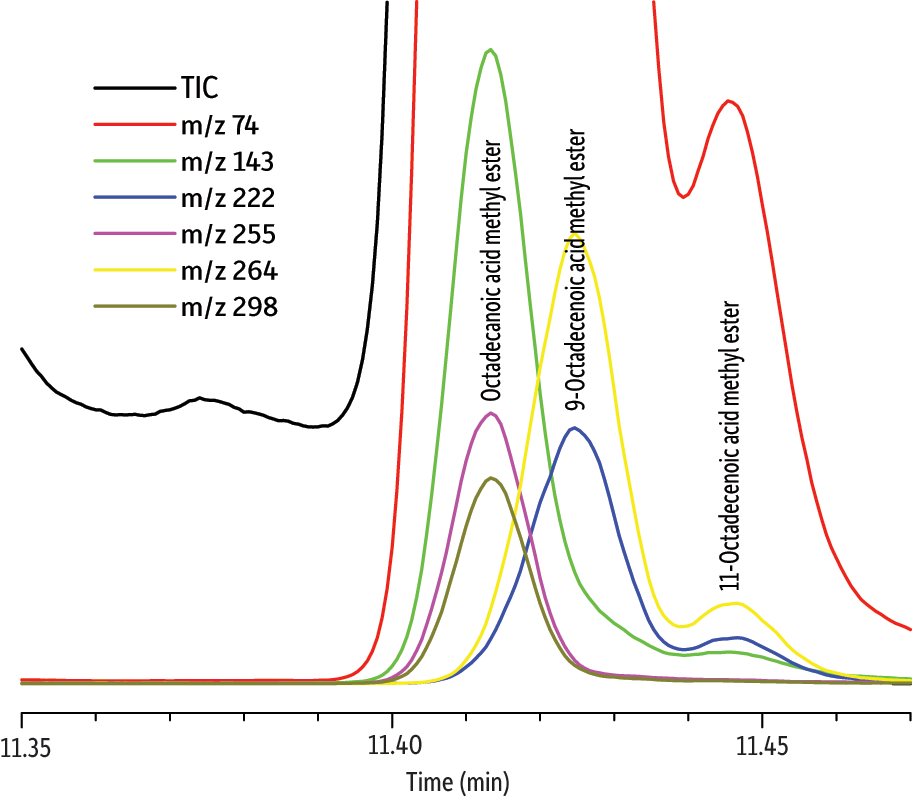
| Peaks | |
|---|---|
| 1. | Octadecanoic acid methyl ester |
| 2. | 9-Octadecenoic acid methyl ester |
| 3. | 11-Octadecenoic acid methyl ester |
See Figure 3 for instrument conditions.
Acknowledgements
The author would like to thank to Restek Corporation for the kind donation of the capillary columns and DANI Instruments for use of the Master GC-TOFMS.
